Average Fragment size
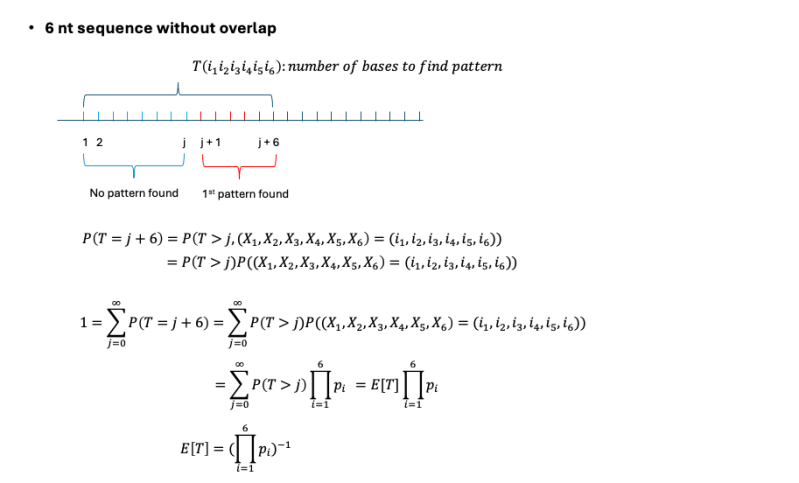
A 6-nt sequence with overlapping occurrences may have a higher total frequency but is expected to have a longer average distance between truly independent occurrences compared to an overlapping sequence. A 6-nt sequence can appear in a longer DNA strand in two different ways: with overlaps (where occurrences share some nucleotides) and without overlaps (where each occurrence is fully independent of the others). If a sequence with overlap, then the overlapped suffix nucleotide position in the DNA has a chance to start a new match. For example, in a sequence like "TAATTA", the suffix overlap sequence is TA, which can initiate a new match if ATTA follows. Below is the derivation of the expected length of bases required to find each pattern.
Simulation
The example sequence with overlaps, the expected fragment size, and the simulated fragment size are shown in the table below. A 1 Mb-long sequence is generated 1000 times, and in each case, the average fragment size is calculated and then averaged over 1000 simulations. The frequency of each nucleotide is identical, with p = 0.25.
| Overlap | Ex. Sequence | Average size [bp] | Expected size [bp] | Simulation 1 Mb sequence x 1000 |
|---|---|---|---|---|
| 0 | GATATC | p^(-6) | 4096 | 4063 |
| 1 | TCCGGT | p^(-6) + p^(-1) | 4100 | 4086 |
| 2 | TAATTA | p^(-6) + p^(-2) | 4112 | 4099 |
| 3 | TCATCA | p^(-6) + p^(-3) | 4160 | 4142 |
| 4 | ATATAT | p^(-6) + p^(-4) + p^(-2) | 4368 | 4341 |
| 5 | AAAAAA | p^(-6) + p^(-5) + p^(-4) + p^(-3) + p^(-2) + p^(-1) | 5460 | 5414 |
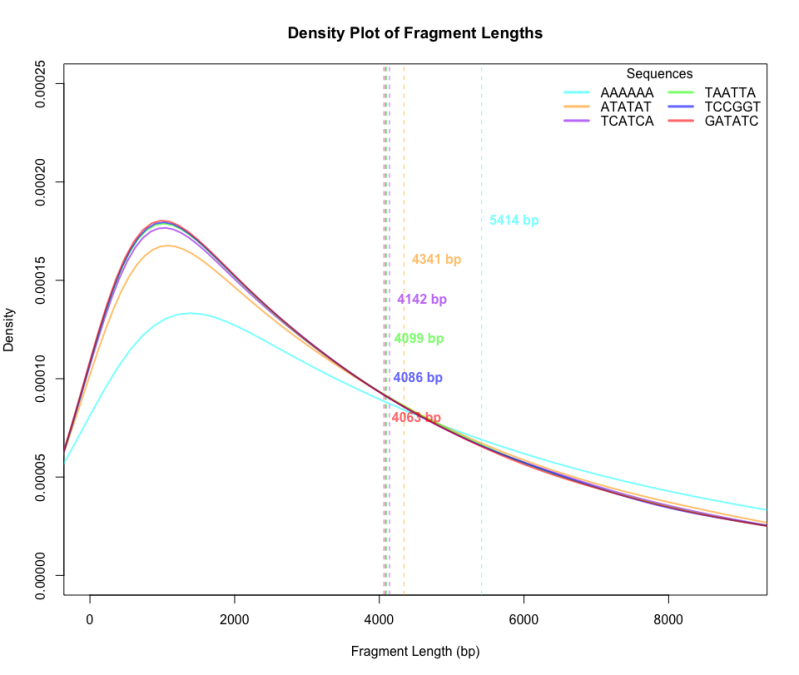
Density plot of fragment lengths generated from 1000 simulations of a 1 Mb DNA sequence. When a 6-nt sequence partially overlaps with itself (by 1 to 5 bases), the average fragment size increases, shifting the distribution to the right.